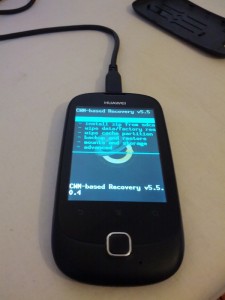
I’ve just finished porting ClockworkMod Recovery to the Huawei U8185 (Ascend Y100). It took a bit longer than expected because I didn’t have a stock ROM to inspect, and I was playing it a bit safe. Long story short: ClockworkMod Recovery is now available for the Huawei U8185.
I’ve tested flashing Superuser as well as a nandroid backup. I haven’t yet tried a restore. There is an issue with the back-button functionality (soft keys don’t work, and the “Go back” menu option doesn’t always appear).
Download
At the time of writing there are binaries available at the following locations:
- clockworkmod_5.5.0.4_u8185.img: Mediafire / BayFiles / goo-inside.me (md5:
85b44d75d9d73bdacc0ecc521ef9259a)
Flash
Put your phone in bootloader mode (yank battery, then start your phone while holding Volume Down for ~5 seconds). Connect your phone to the computer with USB and flash with fastboot1:
fastboot flash recovery clockworkmod_5.5.0.4_u8185.imgBoot into recovery
If you want to boot into recovery mode you can do it at boot, by holding Volume Up for ~5 seconds as you power the device on (make sure you’ve removed the battery first), or you can simply reboot directly to recovery mode using adb:
adb reboot recoverySource
The source code for ClockworkMod Recovery is open source as part of the CyanogenMod source tree. This recovery was built using a custom device tree I wrote for CyanogenMod 9; you can find the source code here: github.com/alanorth/android_device_huawei_u8185
Happy hacking!
1 I’ve uploaded a copy of fastboot for convenience, but you should be able to find a copy somewhere if you just search. It’s an open source tool.
Full swing.Indeed!
Hey Allan,
Quick one, I can flash Clockworkmod without rooting right?
Yep. You just reboot the phone into bootloader mode and then flash the partition with the file I provided. 🙂
hi i need help im stuck the volume down + power button doesn’t work i dunno what happened but i can still go to the volume up+power button now my phone is stuck can you help me please….
i’m stuck on a colored screen when i try to turn on my phone help i just bough my phone …………..
If you tell me the screen is colored, it helps to say which one? Is it pink? Yank the battery out and boot it again.
How long exactly should it take for bootloader mode, and how do I know its in bootloader mode? I am stuck at the first step with the Huawei logo showing, for like 10 minutes…
OMG sorry, dumb question. I apologize profusely, continuing with steps
I have booted into recovery, what next? is the phone rooted yet? Haha..kiddin, I am tryin the commands on the other post, to flash superuser, and am getting “Cant find /system in /proc/mounts” …
hi Allan, am rooted, but got some development. I’ve been trying to create own rom but got one problem, may be you can help. i downloaded a cyanogen custom rom for u8150, copied my system files for u8185 and replaced system files for ub. y 8150, everything properly arranged but cant install, coz the MTN-INF, is still for u8150, maybe you can assist. thanks
You’re trying to install the U8150 CyanogenMod for U8185? I would rather put my time into doing a source port. I’ve already set up a device repository: https://github.com/alanorth/android_device_huawei_u8185. That’s where we should be putting our energy! Also, we should be doing CM9 instead of CM7.
it’s META-INF for zip installation not MTN-INF
hi Alan, i will try to work on cm9 and any progress i will let you know. thanks.
hi Alan, I created a cyanogen rom, on the 1st installation it failed displaying no booting.img, on my second installation displayed installation complete but had a problem on booting, that’s my progress. How i created the rom; i had a cyanogen zip rom, extracted, downloaded root browser, used it to copy system folder( copied file after file, arranging them in a similar folder created in the sd card, eliminated files that had no importance, used the same format as the cyanogen zip file) I created a META-INF file for zip installation which worked properly. hey, was i heading the correct direction?
I dunno. I don’t waste my time doing zip porting of ROMs. I’m going to use source, and create a real one. 😉
Works marvelous! Thanks Allan!
Hi Allan. Great stuff. However am stuck. I’ve successfully flashed clockworkMod but can’t find the superuser zip at the link you provided (http://goo.im/superuser). Can i use any superuser zip e.g the one at http://downloads.androidsu.com/superuser/Superuser-3.0.6-d-signed.zip, or must i have the one u specified?
eeh never mind. Just installed the latest superuser zip from androidsu.com and it worked perfectly! 🙂 My u8185 is now rooted!! thanks
You can’t find the link because goo.im’s servers crashed recently. Twice. So devs haven’t re-uploaded their files yet. Glad you found Superuser somewhere else, though. btw, The “efghi” means “Eclair, Froyo, Gingerbread, Honeycomb, Ice Cream Sandwich”… (Android versions).
Hi Alan, how far now have you gone on CM9 working with U8185?
Hey,
I didn’t have time to post it here yet, but I posted to Google Plus, Twitter, and the Nairobi Linux Users Group. Here’s the status of CyanogenMod 9 on the U8185: https://plus.google.com/101880444013156960674/posts/gn2jgiasgvu (hint: it’s booting, and running like a dream).
are you supposed to copy the clockworkmod_5.5.0.4_u8185.img to your sd card?
No. You have to put your phone in to fastboot mode, and then flash the image to the phone from your computer. As I explained in the blog post 😉
I can not put mi mobile in bootloader mode ,my screen frezze itself .There is other method to do it? thanks and thank you
In bootloader mode the screen is frozen at the Huawei logo. 😛
Thank you Alan I can do it,but now ,do you know if exist a cm for this mobile? thank you and congratulations your blog is great. From Mexico
entro a modo flash pero me apareceré después de teclear fastboot “waiting for device” necesito otros controladores
Marco, try typing
fastboot devices. You should see something like this:That means fastboot can see your device. If it only says “waiting for device” you might need to try
sudo fastboot devices.linuxmint 13
marco@marco ~/Descargas/android-sdk-linux/platform-tools $ ./adb devices
List of devices attached
24DBAC255569 recovery
marco@marco ~/Descargas/android-sdk-linux/platform-tools $ sudo ./fastboot devices
[sudo] password for marco:
marco@marco ~/Descargas/android-sdk-linux/platform-tools $
ok, bootloader power + vol- windows 7 x64 gracias
instalado superuser no root
Hola Humberto. I’m from San Diego, California. I used to eat tacos in Baja California… but now I live in Africa and there are no tacos or carne asada anywhere. 🙁
No, there is no CyanogenMod for this phone… yet. I have a VERY beta CM9 and CM10, but the critical stuff like phone, SMS, 3G, wifi, rotation sensors, etc aren’t working yet. Just keep following me on Twitter and Google Plus, we’ll get CyanogenMod soon… it just takes time to port 😛
It worked great for me on my o2 phone. Got rid of all the bloatware, changed bootanimation.zip (stored in /cust/o2/uk/media/) Cant find the splash logo though 🙁
Thanks!!
Glad it could help you. I still haven’t spent much time on this phone… but I have managed to get CyanogenMod 9 and 10 (ICS and JB) to boot on it, though it still needs a lot of work (modem, wifi, etc). I hate the stock Huawei ROM!
install version 3.1.3 thank’s allam
hi allan. i would really need some hel here. am trying to root my phone n having a problem in flashing the phone with clockwork mod. 1st i downloaded the android sdk pack which has the fastboot (u recommended for flashing)and installed it in local c:. i tried to open fastboot but nothing happened. i opened cmd prompt. typed c: trying to find the path of fastboot but it was not recognised. would someone pliz give me a heads up bcoz i really need a rooted phone. i would really appreciate if someone clearly notes down the steps of installing clockwork mod n superuser for me n other android newbiez.
Sorry, I don’t know how to do it on Windows… I haven’t used Windows in like 10 years 🙁 When I get time I may try to find a way to flash ClockworkMod Recovery from the U8185’s existing stock recovery… no promises though!
Hey guyz…i really need a link or a custom ROM for huawei U8185….i flashed the previous ROM and now am stuck I cannot find a compatible ROM for this device…pliz help.
I don’t think there ARE any custom ROMs for this device… I started porting CyanogenMod 9, but haven’t worked on it much since August. Keep checking my blog, I’ll start back up soon hopefully.
hola e conseguido entrar al recovery pero sin ser root y no estoy muy seguro de meter una rom diganme q puedo aser y como realizar bakud de mi rom actual ya q no se como acerla contesten a mi correo porfa g.29079@hotmail.com
Que tal Gabriel
Lograste rootear tu Huawei Ascend Y100, U8185????
crees que me podiras ayudar a indicarme como hacer con el mio?? tengo problemas de espacio por los programas que vienen de fabrica y el volumen es muy limitado.
no logro entender los foros, mi ingles debe mejorar y no puedo permitirme dañar el cel.
Gracias por tu colaboracion
Atte.
Hey Alan, just rooted my 8185,thank you very much
Awesome! Glad to have helped. One day I’ll write up proper instructions… including an XDA forum post. This phone isn’t as popular in Kenya yet, but I want to make sure the information is ready once the people decide to search for it!
hi Alan.. i am new to this rooting android. i need help with my u8185. i decided to root my device to free up memory usage and to customize it.. i tried but i was stuck with recovery.. some applications were removed like google play and this touch keyboard application.. but still when i tried to install rom manager, it showed that my phone needs to be rooted… i dont know what to do.. i tried to search and understand the steps but still i don’t get it.. im using windows.. is there a more simple way of how to root?for dummies? thanks
Don’t install ROM Manager; it won’t help you on this device. I don’t know how to use Windows, so I’m afraid I can’t help you for now. Once I get time I may find someone to make an easier method for ClockworkMod Recovery, and with Windows-specific instructions.
c you just do a complete video on the whole process because its simpler than written instructions
I don’t have a video camera… but that’s a good idea. One day maybe. 🙂
hey allan….i have a ln to a custom rom http://android.podtwo.com/roms/r0_de-branded_hwu8185-update-signed.zip but it has issues with 3G internet services…could you check it out for me and try to rectify it?…then upload it
I downloaded that one the other day but haven’t tried it yet. I don’t have time to mess with it right now, sorry!
thank you man it work perfect … but when I boot into recovery mode whatever command I choose the CWM display the logo (the hat over the arrow) and not responding what is wrong????
Use the power button to go backwards. Use the menu key to enter, and arrows to select. I will post a new version of CWM Recovery soon (version 6.0.1.x), which is much easier to use.
Hi Alan,
I am not a genius of phone softwares but I am sure with the proper instructions from you I can root my Ascend Y100 that I bought recently.
The best thing you can do now for those of us think all this is rocket science is to give the most detailed of instructions on rooting the Ascend Y100.
Personally I have net yet succeeded in opening the bootloader.
I hope you will hear my cry and respond accordingly.
regards,
Geoffrey.
I wish the original recovery for u8185 due the need to upgrade firmware, any one who did backup? please
Here’s the stock recovery for U8185: http://www.mediafire.com/?rgupc6pi4p7a52i Good luck!
You guy after u boot into recovery explain to me like im a two year old what you step by step kindly ahsante
What are you trying to do? Have you already flashed using
fastboot?hiiii alan i was trying to find the surperuser zip but any that i have find work on the phone can you try to upload yours ? if you have it ?
hey Allan…finished yet porting the CM9 for huawei u8185?
Nope, not yet. Haven’t spent any time on it this week lately… still need help. It’s too much work for me alone, and I’m bored. 😀
Hey allan,could u plizz explain how to flash the file with fastboot
hey Alan, I use windows in flashing the recovery and also superuser into the device. its simple, just need pdanet for android installed in your computer and the fastboot, adb, adbWinApi and adbWinUsbApi files then start command prompt and use fastboot to flash the recovery into the phone when in bootloader mode.
I rooted my huawei ascend y u 8185, Is there any way on how I can go back to my original mode?
Root is “normal mode”… what are you trying to achieve?
flashed your cm9 its stuck on the boot screen
Make sure to wipe data / factory reset when moving from Huawei ROMs to CyanogenMod.
Plz help ‘mount USB storage’ does not work on Clockworkmod 6.0.1.2
Hi Allan, where can i find the original huawei u8185 romrom. I installed the r0_de-branded rom and im getting my phone hang up every now and again. Besides Wi Fi doesnt seem to work for internet connectivity though the phone “sees” and connects to available hotspots/ connections.
Alternatively, do you have a way to fix this. Its baseband 109808. android version 2.3.6
Thanks in advance
Hey, unfortunately the last time I checked there are stock ROM dumps but no way to restore them. On the U8150 you could restore by putting the APPDATA on the sdcard and rebooting holding a certain key combo (up?). From what I saw on the U8185 it doesn’t work…